Enterovirus coxsackievirus A16 2A protease small scale expression and purification protocol
Korvus Wang, michael fairhead, Eleanor Williams
Disclaimer
Research was supported in part by NIAID of the U.S National Institutes of Health under award number U19AI171399. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Abstract
This protocol details the expression and purification of coxsackievirus A16 2A protease construct bearing a N-terminal His-SUMO tag at small scale (<6L).
Picornaviridae Enterovirus coxsackievirus A16 (CVA16)
Attachments
Steps
Abbreviations
CV - column volume, total volume of resin in a column
IMAC - immobilised metal affinity chromatography
FT - flow through
Plasmid Transformation
Transform the coxsackievirus A16 2A protease plasmid into BL21(DE3) and store a glycerol stock of this at -80°C
The coxsackievirus A16 2A protease plasmid encodes the 2A protease with an N-terminal His6-SUMO tag on a kanamycin resistant backbone with a T7 promoter.
Protein expression
Scrape off some of the glycerol stock with a sterile loop and use this to inoculate a 50 mL falcon tube containing 10mL of LB supplemented with 50ug/mL kanamycin. Grow the starter culture at 37°C 4h 0m 0s with 200 rpm shaking.
Use the 10mL starter culture to inoculate 1L Sample(see Materials) supplemented with 50ug/mL kanamycin in a baffled flask. 200rpm
When the OD600 reaches approximately 3.0, lower the temperature and shaker speed to 180rpm and incubate
Harvest the cells by centrifugation at 4000x g,4°C. Discard the supernatant and store the pellet at -80°C .
Protein Purifcation
Lyse cell pellet
Thaw and resuspend the pellet using ~7mL of lysis buffer per g of pellet. Stir gently at Room temperature for 0h 30m 0s to allow lysozyme and benzonase to start breaking down
cell components.
Lyse cells by sonication on ice 0h 0m 4s 0h 0m 12s for a total 'on' time of 0h 7m 0s at 50% amplitude to fully rupture the cells. Ensure pellet remains cold during sonication to prevent overheating.
Centrifuge the lysed cells for 38000x g,4°C to remove insoluble cell debris, and collect the soluble fraction in a bottle 4°C
Perform IMAC to extract target protein from the lysed cell mixture
Dispense 5mL of IMAC resin (Ni Sepharose 6 FF, Cytiva) into a gravity flow column. Rinse resin with ~ 10CV distilled water to remove the storage solution and then ~ 10CV binding buffer to equilibrate the resin.
Resuspend the equilibrated resin with 10 mL of binding buffer and add to the supernatant bottle. Incubate the resin with the soluble fraction for 0h 30m 0s while rotating or otherwise mixing gently at 4°C
Load the resin/soluble fraction mix back onto the gravity flow column, retaining the flow through separately for SDS-PAGE analysis.
Wash the column with 10CV of base buffer, followed by 10CV of wash buffer 1 and 2. Allow wash buffer to pass through completely between washes. This is to remove non-specific, weak binding of contaminant proteins from the resin for a cleaner elution. Collect washes separately for SDS-PAGE analysis.
Elute the protein with 1.5CV of elution buffer.
Repeat step 8.5 one more time, collecting a total of 2 separate elution fractions. This is to ensure maximum retrieval of protein from the resin. The total protein concentration of the elutions are then measured by Nanodrop.
Wash used IMAC resin with 10CV of base buffer, and leave the column submerged in a small amount of base buffer so that the resin is kept moist. This washed IMAC resin will later be reused for reverse IMAC (rIMAC)
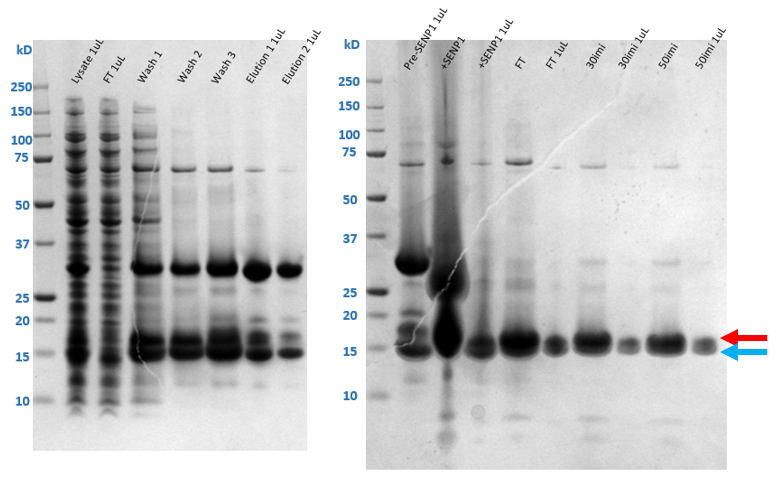
Run SDS-PAGE of all samples from total lysis supernatant to final elution. Stain gel with protein staining solution Coomasssie Blue and determine which fractions contain the target protein by finding the band corresponding to the target molecular weight.
Elution de-salting, tag cleavage and reverse IMAC
Pool the elutions and desalt using a HiPrep 26/10 deasalting column, run on an AKTA pure at a maximum flow rate of 10mL/min.
Add His-SENP1 SUMO protease at a 1:100 ratio to the total protein content of the desalted sample, as determined by nanodrop. Incubate at 4°C This cleaves the affinity tag.
Pour the cleaved Coxsackievirus A16 2A protease, SUMO tag, SENP1 protease mixture over the washed IMAC resin and collect the flow through, rIMAC.
Wash the IMAC resin with 2CVwash buffer 1 and 2 to remove any target protein still bound to the resin. Take samples of the FT and washes for SDS-PAGE analysis.
(Optional) elute rIMAC resin with 2CV elution buffer to confirm if the protein shows non-specific binding to the resin used.
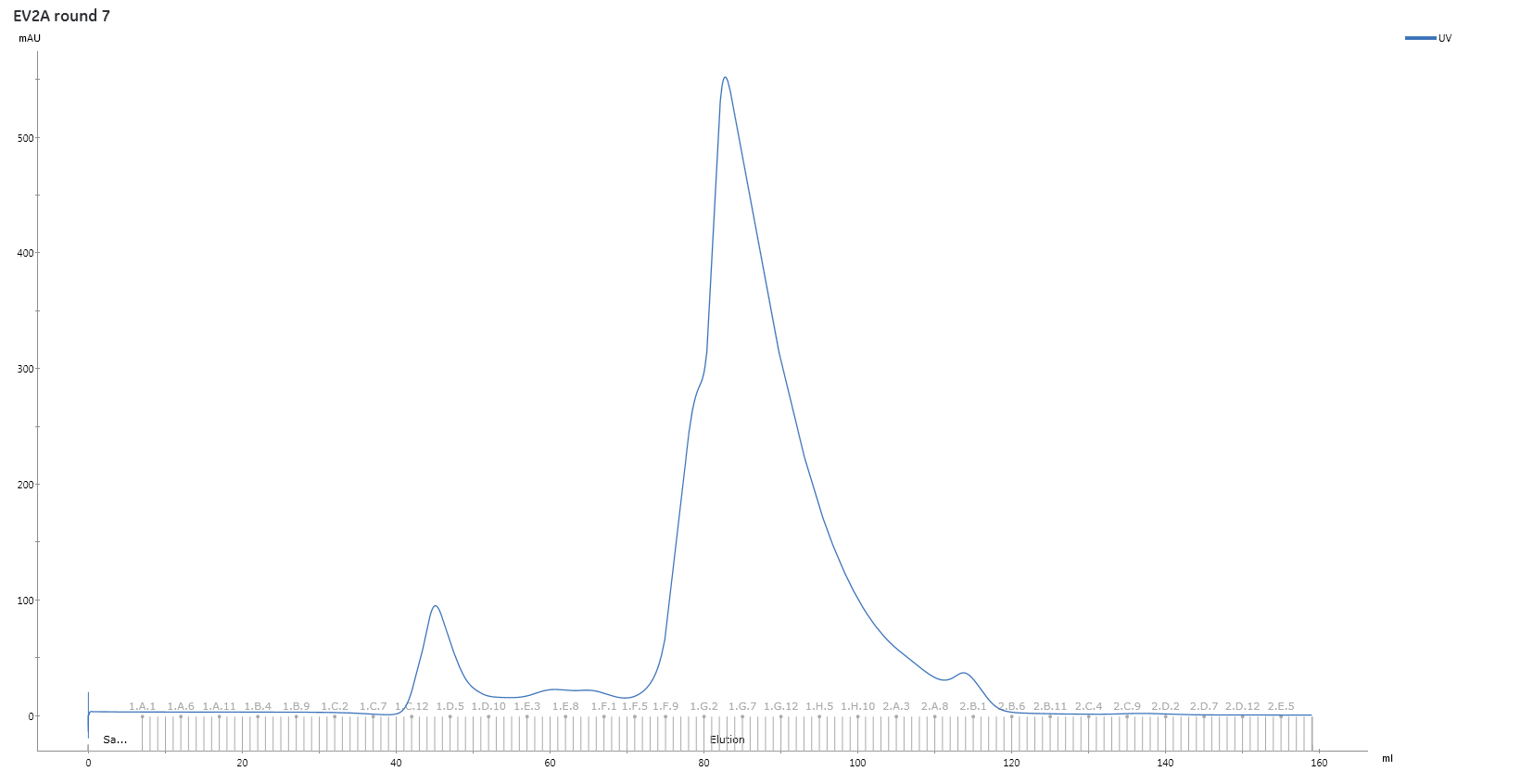
Purify sample further by size exclusion chromatography .
Using 10,000 MWCO spin concentrators, concentrate the rIMAC step containing fractions of the target protein to a final volume of under 5mL .
.
Remove any solid aggregates from the sample by centrifugation at 17200x g,4°C , then immediatly draw up the supernatant with a 5mL syringe and a blunt-tip fill needle, taking care not to disturb the pellet.
Using an AKTA Pure system:
Inject the sample onto a 5mL sample loop and run the sample down HiLoad 16/60 Superdex 75 pg gel filtration column at 1 mL/min using gel filtration buffer as the mobile phase, collect 1mL fractions.
Analyze the size exclusion chromatography fractions by SDS-PAGE and pool the fractions with highest amounts of pure 2A protease.
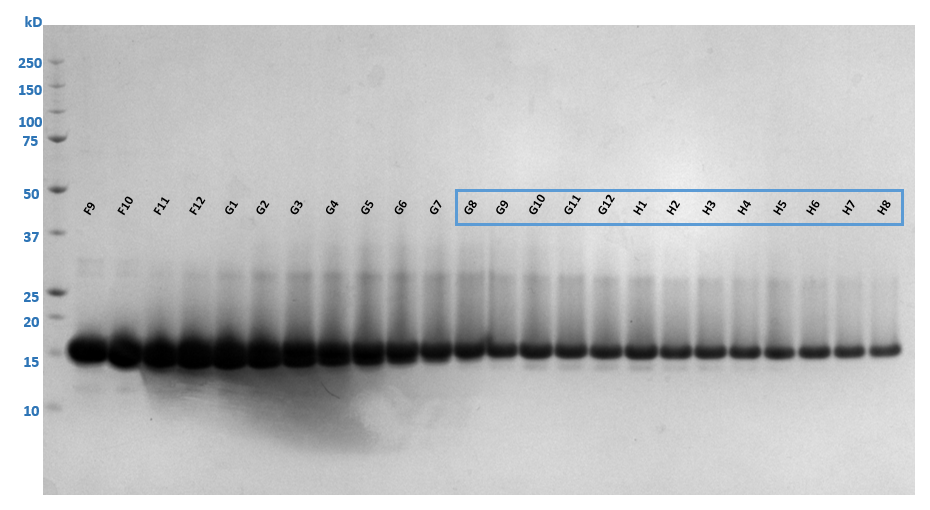
Take the fractions that contain the cleanest target protein and concentrate to 15mg/mL using a 10 kDa MWCO centrifugal concentrator
Take 1µL of the final sample for SDS-PAGE, and another for mass spectroscopy (MS).
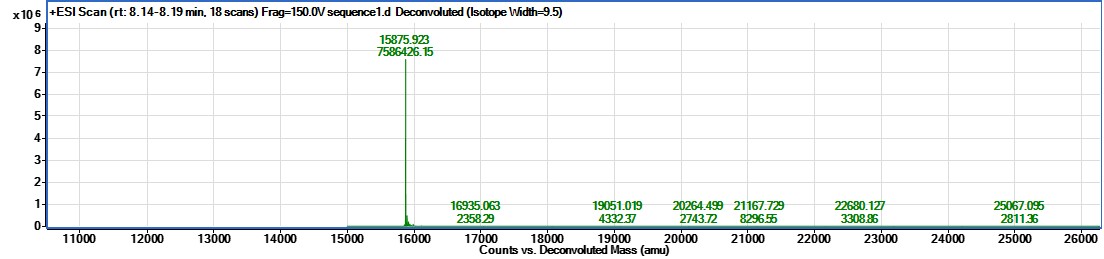
Aliquot into appropriate volumes for future usage to minimise freeze/thaw cycles. Flash-freeze in liquid nitrogen, and store at -80°C until required.
For example:
The final yield from processing 6 L of cells was 50 mg of pure coxsackievirus A16 2A protease

