Concentration and nucleic acid extraction of viruses from wastewater primary sludge
Ari N Machtinger, Olivia S Hershey, William J Bradshaw, Daniel P Rice, Michael R McLaren, Emily H Soice
wastewater-based epidemiology
WBE
viral concentration
metagenomics
viromics
wastewater
wastewater sludge
Abstract
This protocol details our workflow for performing concentration and total nucleic acid extraction from wastewater primary sludge (settled solids) for the purposes of untargeted RNA and DNA sequencing of viruses present in wastewater.
Protocol overview: In this protocol, 20 mL of wastewater sludge is concentrated to a final volume of 400 uL using the InnovaPrep Concentrating Pipette Select. Prior to concentration, the wastewater sample is treated with Tween 20 and sonicated to dissociate viral particles from solids in the wastewater matrix. The sample is then centrifuged to remove larger solids. The pellet is discarded, and the supernatant is filtered with a 0.45 um PES 75 mm filtration unit to remove remaining suspended solids and bacteria. This filtrate is then concentrated with the Concentrating Pipette, using Ultra CPT tips and recommended device settings for the InnovaPrep modified wastewater processing protocol. Nucleic acids are then extracted from the concentrated product using the Zymo quick-DNA/RNA Viral kit using the manufacturer protocol with a few modifications that we have found to be helpful.
Before start
Read the 'Safety Warnings' section for biosafety precautions necessary for handling raw wastewater samples. Prepare the fume hood for wastewater handling, gather materials and reagents (centrifuge tubes, serological pipettes, pipette tips, micropipettes, a marker, strips of parafilm, PBS). Label centrifuge tubes (one for each influent sample, and two for a negative control sample). Ensure proper PPE.
Steps
Reagent Preparation
Prepare a 1% volume Tween 20 stock solution. For a 500mL stock, combine 5mL of Tween 20 with 495mL of 1X PBS. Filter sterilize with a 0.22 um vacuum filtration unit.
Part 1: Sludge Handling, Dissociation, Centrifugation, Filtration
Transfer sludge sample (within secondary container) from the refrigerator to the fume hood. Remove the sample bottle from the secondary container and unseal the bottle by removing the affixed Parafilm.
Prepare aliquots of sludge. Invert the bottle of sludge several times to re-suspend contents, then carefully open it. Using a fresh 50 mL serological pipette, aspirate and dispense 20mL of sludge into a 50 mL centrifuge tube. Repeat for desired number of sample replicates. For a negative control, add 20mL of 1X PBS to a centrifuge tube using a fresh 50 mL serological pipette.
Add 20mL of 1% (v/v) Tween 20 to each centrifuge tube (including controls) for a final concentration of 0.5% (v/v) Tween 20.
Cap and parafilm all bottles and centrifuge tubes.
Place all centrifuge tubes on a vortexer using a 50 mL tube adapter. Shake for 1000rpm
Transfer all centrifuge tubes from the vortexer to a sonication bath, and sonicate for 0h 1m 0s at 40 kHz. Use a paper towel to dry the tubes when finished.
Transfer all centrifuge tubes from the fume hood to a centrifuge equipped with the appropriate rotor and/or adapters (ex: Beckman Coulter Avanti J series with JA-14.50 rotor) . Centrifuge the samples at 10000rpm,4°C. After centrifugation is complete, wait 0h 10m 0s (as recommended by EHS) to allow aerosols to settle before opening the centrifuge. Remove samples from the centrifuge and return them to the fume hood.
In the fume hood, prepare a separate 0.45 um vacuum filtration apparatus for each sample and control by attaching the filtration unit to a clean pyrex bottle.
Decant the supernatant from the sludge sample centrifuge tube directly into a 0.45 um vacuum filtration top attached to a sample collection bottle, taking care not to dislodge the pellet.
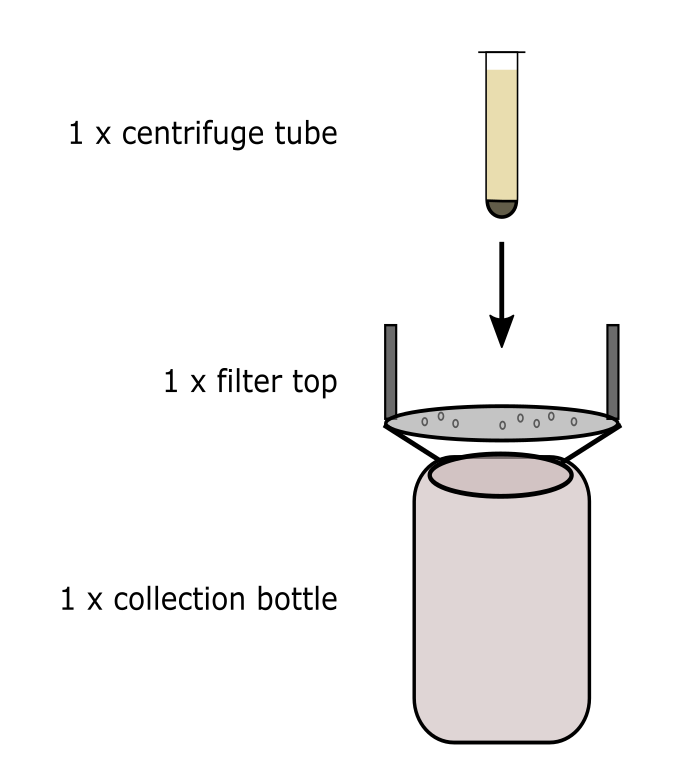
Begin vacuum filtration by capping the vacuum filtration top and opening the vacuum line. When complete, cap the pyrex bottle and set aside. Repeat for each sample replicate.
For the negative control centrifuge tube, decant directly into a 0.45 um vacuum filtration top. There should not be a pellet. Perform vacuum filtration as was done for the sludge, cap the pyrex bottle, and set aside.
Part 2: Concentration via InnovaPrep Concentrating Pipette Select
Perform the "Start-up" protocol for the InnovaPrep Concentrating Pipette Select.
- Turn on the Concentrating Pipette, and navigate to "Maintenance" and then "Start-up". Follow the prompts.
- Check that the maintenance tip is in place.
- Place the waste line in the proper position.
- Remove the storage fluid line and insert the foam elution canister.
- Ensure that the screen reads "WWULTRA".
Run the concentration protocol for the filtered sludge sample (ie, filtrate).
Remove the maintenance tip and place a fresh Ultra CPT into the tip port. Lower the tip into the sample.
Press "Start Run", allow the Concentrating Pipette to aspirate the waste, and wait until it beeps to signal the end of the run.
While holding a 5 mL centrifuge tube under the Tip, press "Elute" and catch the foam that is dispensed.
After the foam degasses, add 400µL of the Zymo DNA/RNA shield reagent.
Run the concentration protocol for the negative control. Perform as is done for the sludge sample.
Incubate samples with added Zymo DNA/RNA shield reagent at room temperature for 0h 30m 0s. If samples are sitting for longer then store at 4°C
Perform the "Shut Down" protocol for the InnovaPrep Concentrating Pipette Select.
- Navigate to "Maintenance" and then "Shut Down".
- Place the maintenance tip into the tip port.
- Remove the elution canister.
- Check to ensure that there is adequate storage fluid and insert the storage fluid line.
- Turn off the device and remove the waste line.
Part 3: Nucleic Acid Extraction - Zymo quick-DNA/RNA Viral Kit
Gather the materials and reagents for the Zymo quick-DNA/RNA Viral Kit in the Biosafety Cabinet. Equilibrate samples to room temperature.
Add 1600µL of Viral DNA/RNA buffer to the sample (a 2:1 ratio) and vortex briefly to mix.
Transfer up to 700µL of lysate into a Zymo-SpinTM IIC-XLR Column in a collection Tube and centrifuge for 12.000x g. Discard the flow-through in the miniprep waste container.
Repeat until full lysate volume is processed.
Add 500µL of Viral Wash Buffer to the column, centrifuge for 12.000x g and discard the flow-through.
Repeat the previous step.
Add 500µL ethanol (95-100%) to the column and centrifuge for 12000x g to ensure complete removal of the wash buffer.
Transfer the column to a clean collection tube, and centrifuge at 12000x g to remove any remaining ETOH.
Carefully, transfer the column into a 1.5 mL nuclease-free tube.
Add 50µL DNase/RNase-Free Water directly to the column matrix and incubate at RT for 0h 1m 0s. Centrifuge for 12000x g to collect the eluate.
Make desired number of aliquots for storage. Place all extracted nucleic acids in a freezer set to -80°C

